-
Filter by category:
- Apps
- Contamination
- Embedded Systems
- Field Solvers
- Interactive
- Mesh Generation
- News
- Numerical Methods
- Papers and Codes
- Plasma Simulation
- Starfish
- Surface Interactions
- Uncategorized
- Visualization
Posts in Category Field Solvers
Multigrid Solver
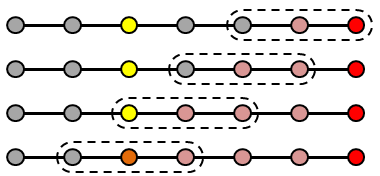
The Multigrid scheme for accelerating convergence of iterative matrix solvers is explained and demonstrated with a one-dimensional Python code. (Mar 11 2018)
Potential Solver for Composite Dielectrics
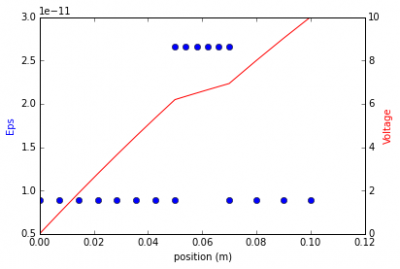
Form of Gauss' law for composite dielectrics is derived. The governing equations are demonstrated with a 1D program that solves electric potential across several pieces with a variable permittivity and mesh spacing. (Jan 18 2015)
Advection Diffusion Crank Nicolson Solver

Online Advection Diffusion equation solver implemented with Javascript and HTML5, and based on Crank Nicolson Finite Volume Method. (Jun 8 2014)
Exporting Magnetic Field from FEMM
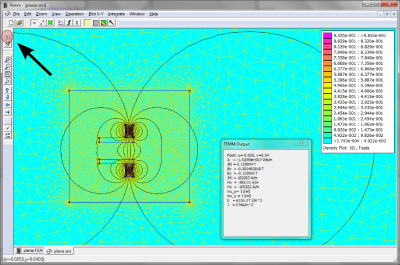
FEMM is a powerful (and free) 2D magnetic, electrostatic, and heat transfer solver. FEMM contains a nice GUI for defining and analyzing the problem. It does not contain an option for exporting the results. However, you can export the results from FEMM easily with a simple LUA script. (Sep 7 2012)
Finite Element Experiments in MATLAB
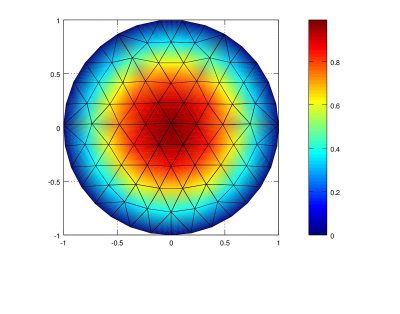
Collection of examples of the Continuous Galerkin Finite Element Method (FEM) implemented in Matlab comparing linear, quadratic, and cubic elements, as well as mesh refinement to solve the Poisson's and Laplace equations over a variety of domains. (Aug 30 2012)
2D Finite Element Method in MATLAB
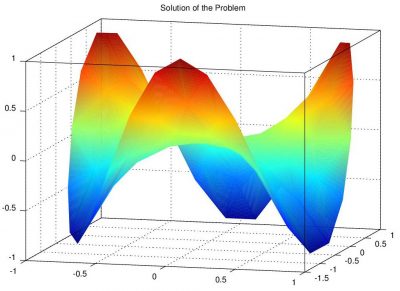
This tutorial discusses how to generate unstructured meshes using a Matlab code distmesh.m and how to use the mesh in a Matlab Finite Element Solver to solve the Poisson's equation on an unstructured domain. (Jun 8 2012)
Nonlinear Poisson Solver
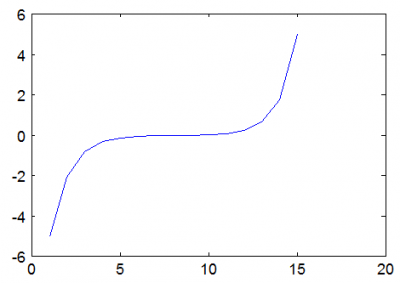
Nonlinear Poisson's equation arises in typical plasma simulations which use a fluid approximation to model electron density. This article describes how to solve the non-linear Poisson's equation using the Newton's method and demonstrates the algorithm with a simple Matlab code. (Apr 30 2012)
The Finite Volume Method
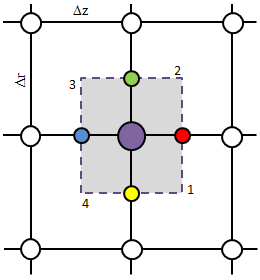
The Finite Volume Method (FVM) is an algorithm for solving differential equations. It is based on the integral formulation of the problem, with each computational element corresponding to a volume over which the integration is performed. It is somewhat more complex than the Finite Difference, however, it can be applied to arbitrarily shaped domains. In this article we show you the basics of this method and provide a simple solver. (Apr 19 2011)
The Finite Difference Method
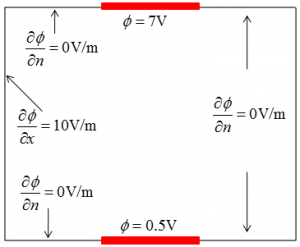
Numerical simulations generally involve solving some differential equations on a computational domain too complicated to solve analytically. The Finite Difference (FD) is a method for solving differential equations. It is a very popular method since it is very easy to understand and implement. Here we introduce the basics of this method and provide a simple example. (Mar 1 2011)